Your pan-seared halibut with lemon caper sauce might not be what you think.
That’s because seafood fraud — due to the intentional or unintentional mislabeling of fish — occurs in at least 20% of species eaten every year in the United States, according to the National Oceanic and Atmospheric Administration.
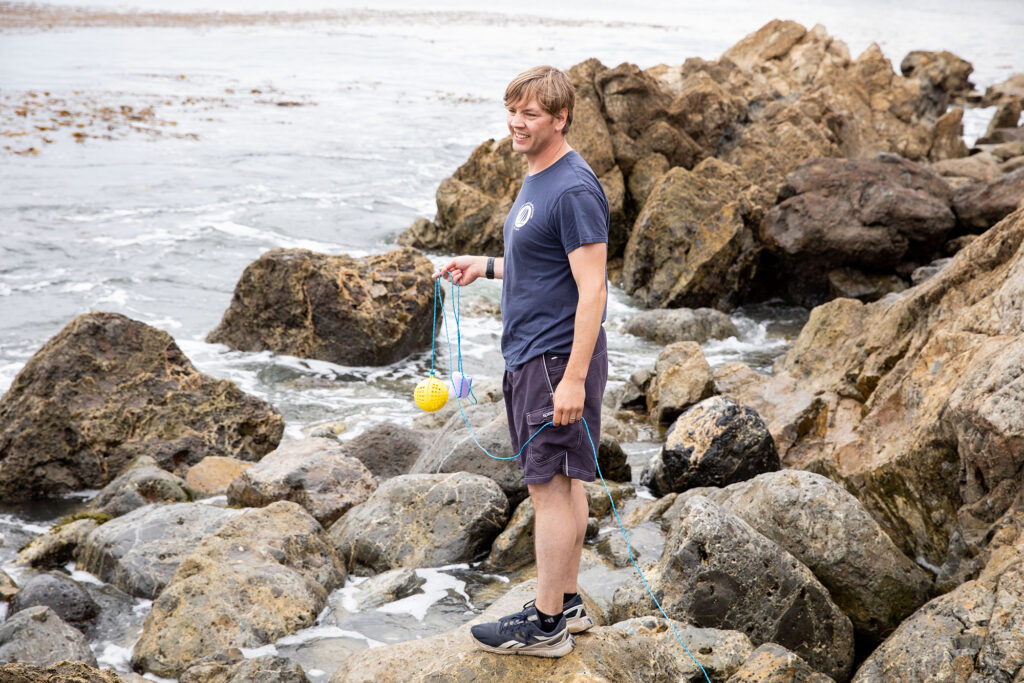
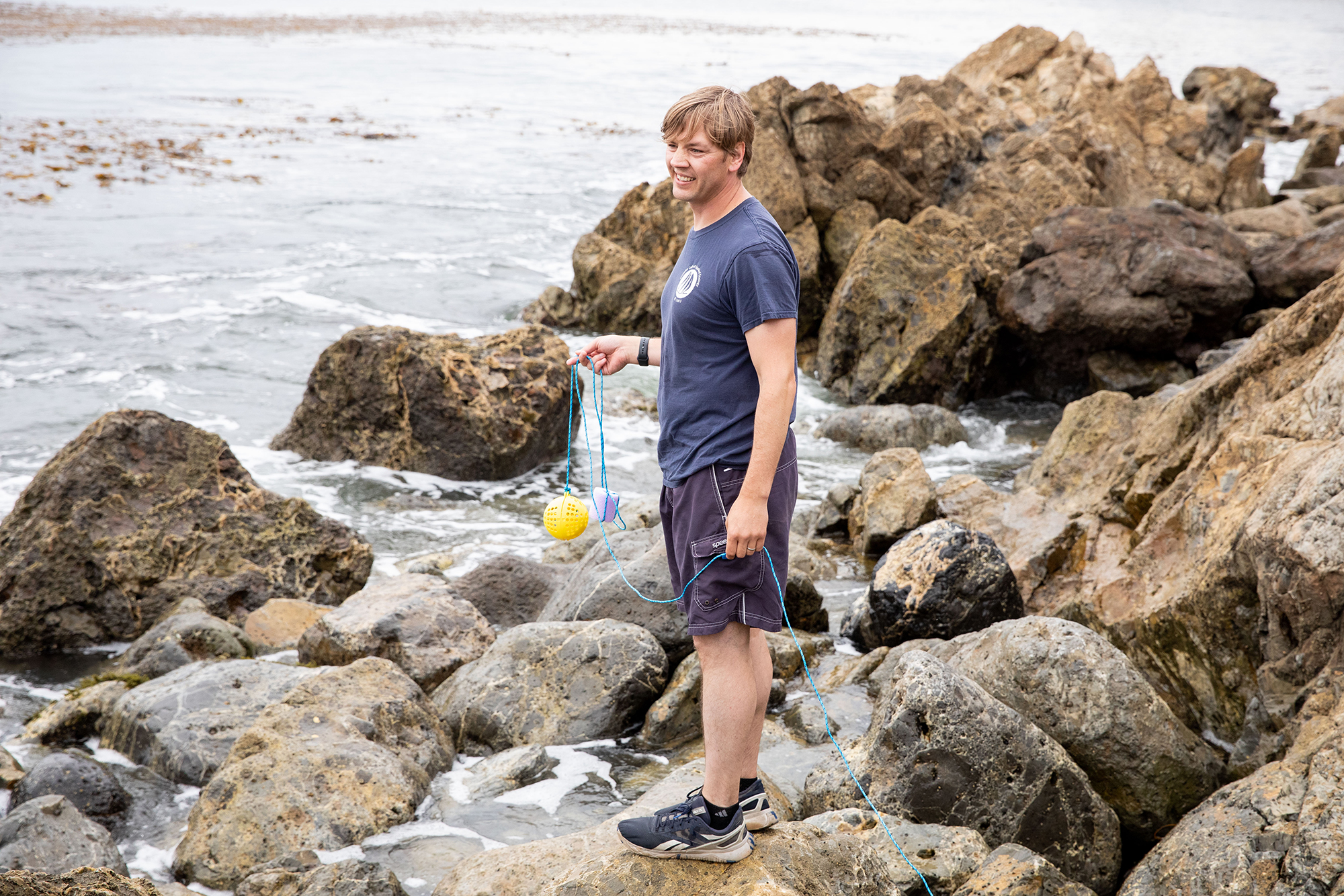
Demian Willette, professor of biology, is turning to eDNA, or environmental DNA, to encourage accurate labeling, and help halt fraud. Willette has been at it for more than two decades, and he’s developed a novel eDNA collection device that could turn the tide on illegal fishing. His as-yet unnamed device gathers scales and metabolic waste from fishing sites and processing plants, allowing researchers to identify fish species through gene sequencing.
The plastic, softball-sized tool, for which he holds a patent, comes in pyramid and spherical shapes, and is dangled in fishing waters or dipped into fish processing holds of ships. Scales and metabolic waste enter the porous collector and lodge on a paper filter. Willette then tests materials and identifies fish species through gene sequencing.
Fish laundering results in an annual economic loss of $83 billion, Willette says.
He developed the device at his startup company, Species Tracks, in collaboration with the biology department of the LMU Frank R. Seaver College of Science and Engineering. Willette says he has a pending contract with the Food and Drug Administration to test the device, which can be manufactured on a 3D printer.
Fish laundering results in an annual economic loss of $83 billion, Willette says. Government regulation is considered essential toward curbing fraud, but scientists like Willette are doing their part to shape the discussion. “eDNA won’t solve this, but it will help,” he acknowledges.
“The end game is, can we accurately trace seafood all the way through the supply chain?” Willette says. “Are you, the consumer, getting what you paid for, and is it safe? In the United States, the burden of proof for validating the identity of seafood is on the processor and the importer. It’s not the fisherman at the beginning of the supply chain, or the restaurant at the end of the supply chain. So we’ve focused our work on the processors.”
Willette’s eDNA collector serves other purposes. It was used by scientists last year off the shores of Yucatan, Mexico, where they detected the illegal use of horseshoe crabs as octopus bait. The Mexican government forbids the practice. Willette’s work not only aids in the preservation of endangered marine species, but it also helps safeguard the bait used to catch them.
Working with the Mexican nonprofit group Comunidad y Bioversidad (COBI), which promotes marine conservation and sustainable fisheries, Willette identified the presence of horseshoe crab DNA in 13 of 15 bait samples obtained from bait containers in ports along the Gulf of Mexico.
Closer to home, Willette and student researchers visited several Los Angeles fish processors last year to further test the eDNA collector. The team did not turn up any problems, but neither was that their motivation. “The work with the processors had been largely in developing the accuracy and effectiveness of the tool,” Willette says. “We have not shared any results from the work with the processors.” At each plant, processers welcomed the researchers.
“I’ve had calls with folks who run processing plants across the United States, and they’re incredibly supportive,” says Willette, adding that that are 600 plants throughout the country. “They absolutely want to see this work, because they understand the potential this has to help their industry become more sustainable. And they want to do the right thing.”
The FDA is responsible for ensuring that domestic and imported seafood is safe, sanitary, and honestly labeled. But it only screens 1% to 2% of all seafood processed, relying on “paper trails,” not hard science, to assess catches, Willette notes. While Willette is a leader in using eDNA to analyze fish, gene sequencing is still considered a developing technology in the battle against fish fraud.
“The end game is, can we accurately trace seafood all the way through the supply chain?” Willette says.
Willette has used students in the fight since 2017. That year, while teaching a marine biology course at UCLA, he got the idea to have students buy seafood from local restaurants, and then sequence their purchases in the lab. Half of the species were mislabeled, they discovered. For four years straight, the project bore similar results.
Willette received a Fulbright Global Scholar Award in 2017, which allowed him to travel to the Philippines and Ecuador, where he first collected eDNA samples from fishing boats. The Philippines, in particular, is a top provider of seafood across the globe.
“I’m a conservation biologist, and what I teach in my classes is how to protect species, while living on this planet alongside them,” Willette says. “Much of that involves first knowing, ‘What are those species?’ You can’t protect them if you don’t know what they are or where they are. eDNA is a tool that helps us get there. It’s a tool that allows us to see the environment as it is, and then it allows us to take action to protect the environment. There are a lot of reasons to be doing this work.”
Andrew Faught is a California-based freelance writer who writes about higher education. His work has appeared in university magazines published by Occidental College, Dartmouth College, the University of California, Berkeley, and the University of North Carolina.

